Overview#
KSTAR Algorithm#
KSTAR is a Python package for inferring kinase activities from phosphoproteomic data in a bias- and error-aware manner, using a graph-based and statistical approach.
In brief, KSTAR works by generating an ensemble of possible kinase-substrate networks based on weighted kinase-substrate predictions (by default, NetworKIN) and calculating the statistical enrichment of a kinase’s substrates in an experiment within each network. The final activity score is obtained by comparing the enrichment of the real experiment to the enrichment that would be expected from a random phosphoproteomic experiment.
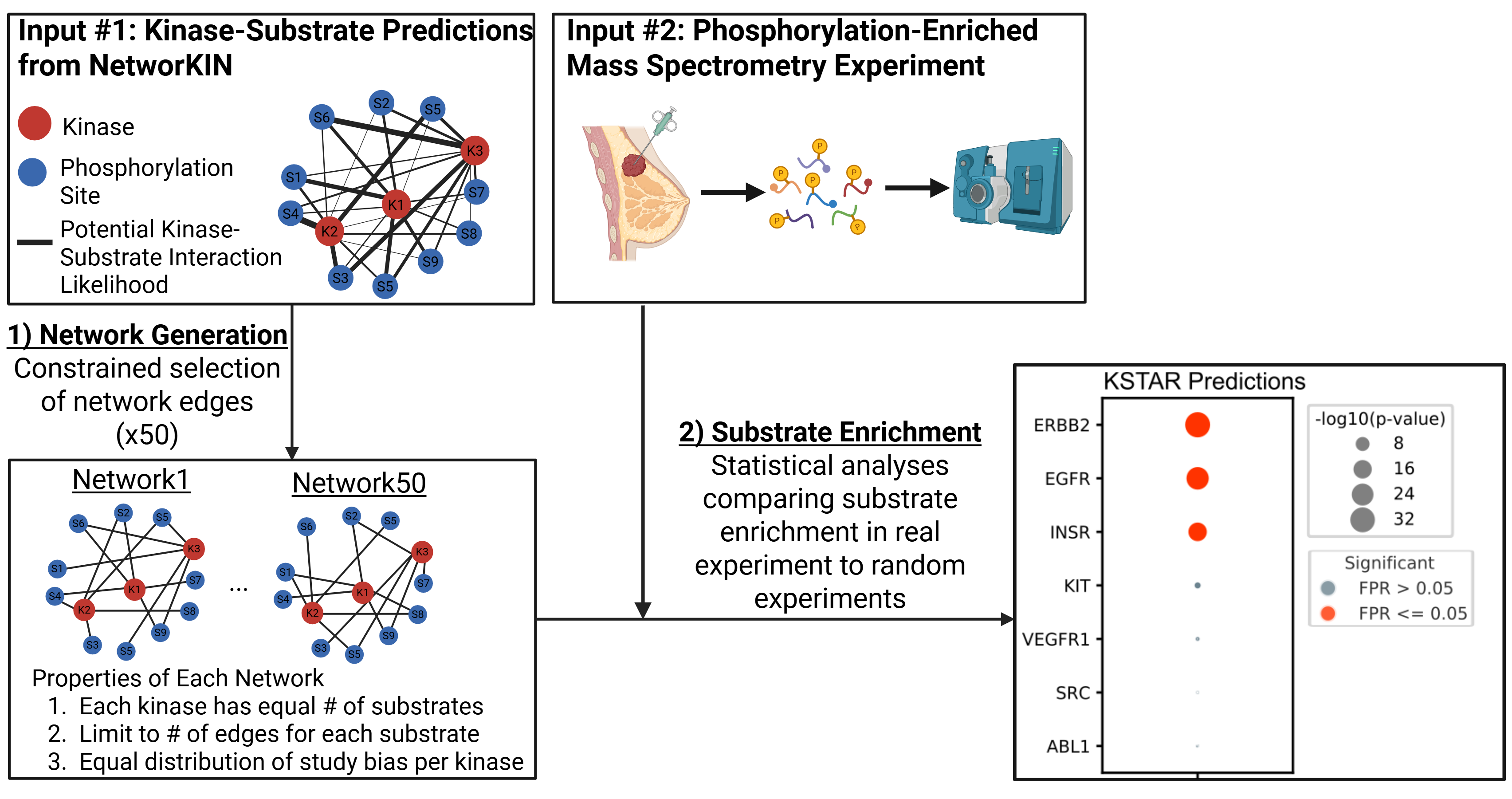
For more details about the algorithm, as well as best use cases, see the original publication: KSTAR Paper
Steps of a KSTAR Run#
For most users, a KSTAR run will consist of the following steps:
Mapping Dataset to Reference Phosphoproteome: Given that the proteome is regularly updated, this step maps phosphopeptides identified in the dataset to the reference phosphoproteome to ensure site positions match the kinase-substrate networks.
Kinase Activity Calculation: Given sites identified in an experiment, for each experiment (column) in a dataset, calculate the likelihood that those sites were pulled randomly from the kinase-substrate networks.
Analysis/Plotting: Various plotting and analysis functions are provided to visualize and interpret KSTAR results.
Note
By default, a pregenerated human kinase-substrate network ensemble based on NetworKIN predictions will be used. However, KSTAR is flexible and can be used with any kinase-substrate prediction graph, not just NetworKIN. If you would like to use your own networks, you can follow the Network Generation tutorial to generate your own network ensemble for use with KSTAR.