Create resampling of available replicate data to explore effects of noise¶
This is similar to the example in Sloutsky, R., Jimenez, N., Swamidass, S. J. & Naegle, K. M. Accounting for noise when clustering biological data. Brief. Bioinform. 14, 423–36 (2013) – we resample each data point according to a Guassian distribution defined by the mean and standard deviation of the measured data.
This example is phosphoproteomic dataset of epithelial cells stimulated with a growth factor and measured immediately afterward at seven time points in the first 32 minutes, specifically [0, 1, 2, 4, 8, 16, 32]. It was downloaded from ProteomeScout and published in: Wolf-Yadlin, A., Hautaniemi, S., Lauffenburger, D. A. & White, F. M. Multiple reaction monitoring for robust quantitative proteomic analysis of cellular signaling networks. Proc. Natl. Acad. Sci. U. S. A. 104, 5860–5 (2007).
Compare this to the example with parallelization
[2]:
import pandas as pd
import matplotlib.pyplot as plt
import openensembles as oe
import numpy as np
fileName = 'experiment.1381.annotated_misTrypsRemoved.txt'
df = pd.read_csv(fileName, sep='\t')
#keep the data and the standard deviations in different dataframes and then instantiate data objects with resampling,
# ultimately merging them together.
dataCols = ['average:data:time(min):0', 'average:data:time(min):1', 'average:data:time(min):2',
'average:data:time(min):4', 'average:data:time(min):8',
'average:data:time(min):16', 'average:data:time(min):32']
stdevCols = ['average:stddev:time(min):0', 'average:stddev:time(min):1',
'average:stddev:time(min):2', 'average:stddev:time(min):4',
'average:stddev:time(min):8', 'average:stddev:time(min):16',
'average:stddev:time(min):32']
descriptiveCols = ['gene', 'mod_sites'] #we don't need ALL the metadata, so we'll just start with this
colsToKeep = descriptiveCols + dataCols
master_df = df[colsToKeep]
x = [0, 1, 2, 4, 8, 16, 32]
stdev_df = df[stdevCols]
stdev_df = stdev_df[stdevCols].astype(float)
#This is the data object with data exactly as it stands with means. We'll create
# a list of new data objects that have resampled data representations
d = oe.data(master_df, x)
[4]:
#create new data objects, resampling each time point for each object according to their mean and stdev
# using a normal distribution
numRepeats = 500
d_arr = []
for i in range(0, numRepeats):
df_temp = master_df.copy()
#get a series from data and make new data column
for index, row in df_temp.iterrows():
for data in dataCols:
stdevCol = data.replace('data', 'stddev')
if stdev_df.loc[index,stdevCol] > 0:
df_temp.loc[index, data] = np.random.normal(row[data], stdev_df.loc[index,stdevCol])
else:
df_temp.loc[index,data] = row[data]
d_temp = oe.data(df_temp, x)
d_arr.append(d_temp)
Cluster all resampled data for an ensemble solution¶
[5]:
transdict = d.merge(d_arr) #merge all data matrices together (could also just iterate through )
[6]:
K = 10
c = oe.cluster(d)
for name in d.D.keys():
c.cluster(name, 'kmeans', 'kmeans_'+name, K)
[7]:
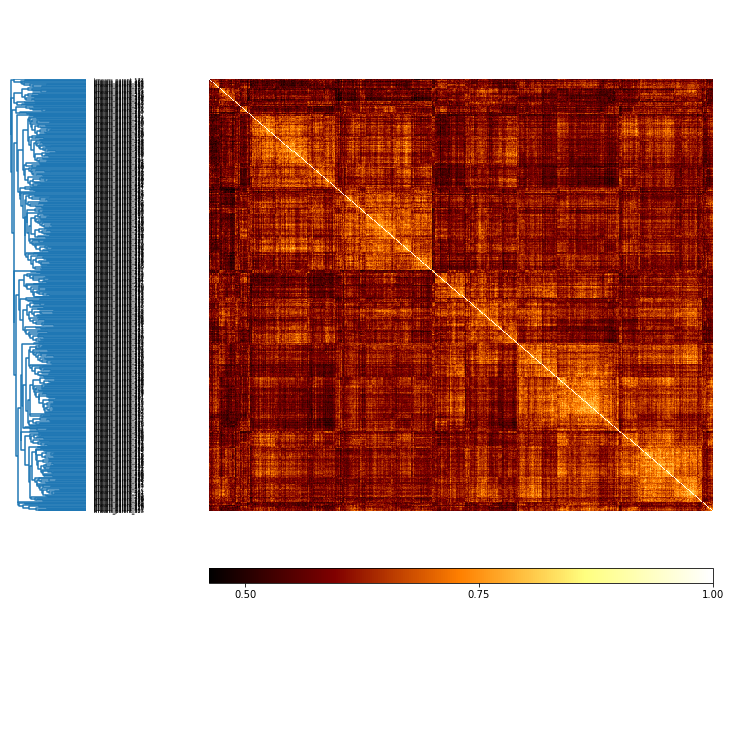
# how do all solutions compare?
MI = c.MI(MI_type='normalized')
mi_plot = MI.plot()
[8]:
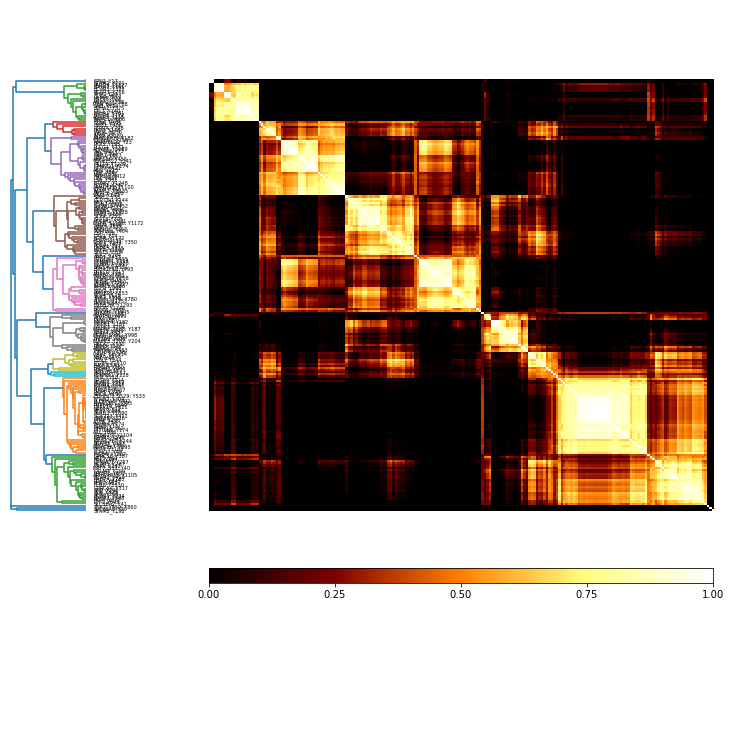
coMat = c.co_occurrence_matrix('parent')
[9]:
labels = d.df['gene'] + '_' + d.df['mod_sites']
co_plot = coMat.plot(threshold=0.5, label_vec = list(labels))